5.2.01. optimising massing

DESCRIPTION
WALLACEI INTERFACE
1. One gene pool component is used to control the uv coordinates on a base surface and another to control the extrusion of floors. Note that gene inputs must renamed with the prefix “wlc”
2. The genes create a randomly extruded massing of three triangular blocks
3. The cast shadow is of the massing is created with occlusion component
4. First Objective is defined as controlling the plot ratio in between 0.1-0.2, a subtraction allows controlled output in a certain range
5. Second Objective is defined as maximising the total floor area, a 1/x component is used to allow Wallacei to maximise the parameter
6. Third Objective is defined as maximising the cast shadow. Note that objectives must named with the prefix “wlc”
7. Phenotypes should be disable during the simulation for a faster simulation. They are only require when results are exported in the Wallacei selection tab
8. The Wallacei X component runs the simulation. Double click to open the interface
9. Exported results are distributed to a grid with phenotypes decoded for further analysis
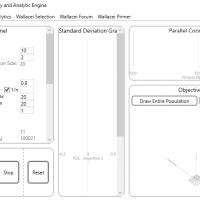
WALLACEI SETTINGS
1.Primary Buttons to Start, Stop and Reset the simulation
2.Basic parameters to control the population size and Mutation : Population:
- Set the generation size (how many individuals per generation) and generation count (how many generations in the simulation) for your simulation.
- Algorithm Parameters: Crossover Probability (0.0 to 1.0) – the percentage of solutions in the generation that will reproduce for the next generation. Mutation Probability (0.0 to 1.0) – The percentage of mutations taking place in the generation (recommends the mutation probability to be 1/n, where ‘n’ is the number of variables (sliders) in the design problem. As such, the default value is 1/n). Crossover and Mutation Distribution Index (0 to 100) – A large distribution index value gives a higher probability for creating offspring near parent solutions and a small distribution index value allows distant solutions to be selected as children solutions.
3.Monitoring the simulation progress, It’s best to optimise your definition if you received “null” results shown in Wallacei
4.Monitoring the SD charts “live” during the simulation
5.Monitor the PCP for the first to last generation “live” during the simulation (red to blue)
6.Monitor the Objective space “live” during the simulation “
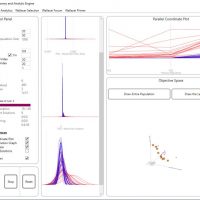
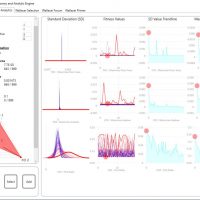
WALLACEI ANALYTICS
1.Primary Buttons to draw the Analytic Graphs, Select Solutions, and Add Solutions to Export List (on tab 3)
2.Select solutions either by name or rank in fitness objective
3.Diamond chart to visualise the ranking of the selected solution
4.Draw the Standard Deviation Charts for all the solutions in the population
5.Draw the Fitness Value Charts for all the solutions in the population
6.Draw the Standard Deviation Trendline for all generations in the population
7.Draw the Mean Value Trendline for all generations in the population
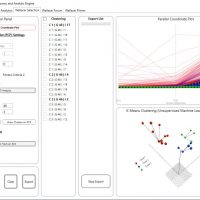
WALLACEI SELECTION
1.Primary Buttons to add solutions to Export list, clear export list and Export phenotypes to the GH canvas
2.Run the 4 analysis methods available in the ‘Wallacei Analytics’ PCP component
a.Most repeated fitness values
b.Solutions with most repeated fitness values
c.Relative difference between fitness ranks
d.Average fitness rank
3.Run K-means Clustering on any generation in the population and add clustered solutions to ‘Export List’
4.Select Pareto Front solutions from the last generation to add to ‘Export List’
5.Displays the solutions associated with the analysis method conducted
6.The phenotypes of the solutions in this list will be exported to the GH canvas
7.Visualise the parallel coordinate plot for all the solutions in the population
8.Visualise the result of the clustering algorithm
This exercise is using Grasshopper version 1.0.0007
Reference: Wallacei(by Wallacei), https://www.food4rhino.com/app/wallacei-0, Accessed August 6, 2020.